Modeling a Pandemic like Ebola with the Wolfram Language
Data is critical for an objective outlook, but bare data is not a forecast. Scientific models are necessary to predict pandemics, terrorist attacks, natural disasters, market crashes, and other complex aspects of our world. One of the tools for combating the ongoing and tragic Ebola outbreak is to make computer models of the virus’s possible spread. By understanding where and how quickly the outbreak is likely to appear, policy makers can put into place effective measures to slow transmissions and ultimately bring the epidemic to a halt. Our goal here is to show how to set up a mathematical model that depicts a global spread of a pandemic, using real-world data. The model would apply to any pandemic, but we will sometimes mention and use current Ebola outbreak data to put the simulation into perspective. The results should not be taken as a realistic quantitative projection of current Ebola pandemic.

To guide us through the computational science of pandemics, I have reached out to Dr. Marco Thiel, who was already describing various Ebola models on Wolfram Community (where readers could join the open discussion). We have worked with him to code the global pandemic model below, a task made considerably easier by many of the new features recently added to the Wolfram Language. Marco is an applied mathematician with training in theoretical physics and dynamical systems. His research was featured on BBC News, and due to its applied mathematical nature, concerns very diverse subjects, from the stability of our solar system to patterns in the mating behavior of fireflies to forensic mathematics, and much more. Dealing with this diversity of real-world problems, Marco and his colleagues and students at the University of Aberdeen have made Wolfram technologies part of their daily lives. For example, the core of code for this blog entry was written by India Bruckner, a very bright young pupil from Aberdeen’s St Margaret’s School for Girls, with whom Marco had a summer project.
The current Ebola outbreak “is the deadliest, eclipsing an outbreak in 1976, the year the virus was discovered,” according to The New York Times. Its data summary as of October 27, 2014, states that there are at least 18 Ebola patients who have been treated or are being treated in Europe and America, mostly health and aid workers who contracted the virus in West Africa and traveled to their home countries for treatment. The C.D.C. reported in September that a worst-case scenario could exceed a million Ebola cases in four months. There are no FDA-approved drugs or vaccines to defend against the virus, which is fatal in 60 to 90 percent of cases and spreads via contact with infected bodily fluids. Here is the current situation in West Africa in the pandemic locus, according to the numbers from The New York Times:


Data Source: The New York Times
Vitaliy: Marco, do you think mathematical modeling can help stop pandemics?
Marco: The recent outbreak of the Ebola virus disease (EVD) has shown how quickly diseases can spread in human populations. This threat is, of course, not limited to EVD; there are many pathogens, such as various types of influenza (H5N1, H7N9, etc.) with the potential to cause a pandemic. Therefore, mathematical modeling of the transmission pathways becomes ever more important. Health officials need to make decisions as to how to counter the threat. There are a large number of scientific publications on the subject, such as the recent Science publication by Dirk Brockmann, which is available here. Professor Brockmann also produced videos to illustrate the research, which can be found on YouTube (video1, video2, video3). It would be interesting to reproduce some of the results from that paper and generally explore the subject with Mathematica.
Vitaliy: How does one set up a computational model of a spreading disease?
Marco: Detailed online models, such as GLEAMviz, are available and can be run by everyone interested in the subject. That particular model contains, just like many other similar models, three main layers: (1) an epidemic model that describes the transmission of the disease in a hypothetical, homogeneous population; (2) population data, that is, distribution of people and population densities; and (3) a mobility layer that describes how people move. I used a similar model that uses the powerful algorithms of Mathematica, its built-in databases, and its powerful data import capabilities. Also, Mathematica‘s visualization algorithms allowed for developing a new model for the spreading of diseases. Some advantages of a DIY model are that we fully control the program and can amend it to our requirements. Mathematica‘s curated data is a very useful modeling starting point and can be paired with flexible data import from external web sources. This is one application of Conrad Wolfram’s idea of publicly available computable data. Using the current capabilities of Mathematica, we can get a glimpse of what will be possible when governmental/public data becomes computable.
There are many different types of epidemic models. In what follows, we will mainly deal with the so-called Susceptible Infected Recovered (SIR) model. It models a population that consists of three compartments: susceptibles can become infected upon contact with the infected; the infected do recover at a certain rate.

To model the outbreak with the Wolfram Language, we need equations describing the number of people in each of these categories as functions of time. We will first use time-discrete equations. If we suppose first that there are only three categories and no interaction between them, we could get the following:

This means that the number, actually the percentage, of Susceptibles/Infected/Recovered at time t+1 is the same as at time t. Let’s assume that a random contact of an infected with a susceptible leads to a new infection with probability b; the probability of a random encounter is proportional to the number of susceptibles (Sus) and also to the number of infected (Inf). This assumption means that people are taken out of the compartment of the susceptibles and go into the infected category.

Next, we assume that people recover with a probability c; the recovery is proportional to the sick people; that is, the more who are sick, the more who recover.

We also need initial values for the percentages of people in the respective categories. Note that the “interaction terms” on the right-hand side always add up to zero, so that the overall population size does not change. If we start at initial conditions that add up to one, the population size will always stay one. This is an important feature of the model. Every person has to stay in one of the three compartments; we will take great care to make sure that this is also true for the SIR model on the network that we describe later! There is, however, some flexibility of how we can interpret the three compartments. In our final example we will, for example, consider deaths. It might seem logical to think that these “leave” our population. In order to keep our population constant, which is important for our model, we will then use a simple trick: we will interpret the last group, the Recovered (Rec), as a set that contains the truly recovered and the dead. It is a reasonable assumption that neither the dead nor the recovered infect other people, so they are inert to our model. Our simple assumption will be that a fixed percentage of people of the Rec group will be alive and the remainder will be dead. Hence, we include dead people in our model—so that they don’t actually leave the groups—and we do not consider births. This results in a constant population size.
This is a naive implementation of the SIR model, which allows you to change the parameters:
We use vectors Sus, Inf, and Rec and iterate them. We will later develop a more direct implementation. Note that the parameters b and c “parametrize” many effects that are at this stage not directly modeled. For example, the infection rate b does describe the risk of infection and therefore models things like population density (high density might lead to more infections) and behavior of people (if there are many mass events, that might increase the infection probability—so does schooling!). The recovery rate c might describe things like quality of the health care system, availability of physicians, and so on. Later we will try to model some of these effects more directly.
The SIR model might not be the most suitable to describe an Ebola outbreak. It is, however, not too far off either. People get infected by contact; the Recovered category might be interpreted as holding the percentage of people who have either survived or died, if we assume that reinfection is unlikely. In different countries/circumstances, the recovery/death rate might vary substantially—something we will model later explicitly.
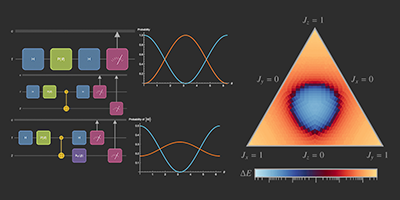
A more systematic way of looking at the overall behavior of the SIR model is to study the so-called parameter space. We can represent how different characteristics, like the highest number of infected or the total number of people who get infected in the course of the outbreak, depend on the parameters. The axes of the following diagram show the infection and recovery rates, and the percentage of people who contract the disease during the outbreak is color-coded:

This shows that for small recovery rates and large infection rates, more than 90% of people contract the disease, whereas for large recovery rates and low infection rates, the total percentage of infected is about 5%, which, in fact, equals the initial percentage of infected.
Vitaliy: To go from pure mathematical to real-world simulations, we would need data, such as populations and their geographic locations. How could data be accessed?
Marco: We will later couple different subpopulations (e.g. airports, cities, countries, etc.) and study the spreading of the disease among these. Each subpopulation is described by an SIR model. When we start coupling the subpopulations, their individual sizes will play a crucial role. Population data, like many other types of data, is built right into Mathematica, so it is quite easy to use that for our modeling.
We will use built-in data to improve our model toward the end, but for a start we could use the international network of all airports to model the transport of the disease. We first need a list of all airports and all flight connections. On the website openflights.org you will find all the data we need. I saved the file “airports.dat” and the file “routes.dat.”
Vitaliy: We could use the latest Semantic Data Import feature to interface with external data. SemanticImport can import a file semantically to return a Dataset object. Dataset and Association are new functions and represent a structured database based on a hierarchy of lists and associations. Dataset can represent not only full rectangular multidimensional arrays of data, but also arbitrary tree structures, corresponding to data with arbitrary hierarchy. It is very easy to access and process data with Dataset, and we will make use of it. A tiny fraction of airports.dat was cleaned up for more precise semantic import. All data files used here are attached at the end of this post, together with the Mathematica notebook.
Marco: Yes, indeed. SemanticImport is very powerful. In my first modeling attempt I used Import and then the new Interpreter function, both of which are very powerful, too. I then needed to do some more “cleaning up” of the data; a multistep procedure that is a common problem when you use external non-computable data. But thanks to your suggestion to use SemanticImport, I could make the code much more concise and readable:

Vitaliy: Yellow-framed entries are semantically processed as Entity:

So we notice that SemanticImport automatically classified the third and fourth columns as cities and countries and converted them to Entity, which is the built-in data representation in the Wolfram Language.
Marco: We can now plot all airports worldwide.
Vitaliy: Indeed, with the new functionality GeoGraphics and its numerous options such as GeoBackground, GeoProjection, and GeoRange, we can tune up the image to a balanced representation of a massive amount of data:

The fifth column in airports is a three-letter IATA airport code. We will need this airport identification code because it identifies connecting routes between airports in the second dataset. Not all data entries have it; for example, here are the last 100 cases:

Some of these are also false because they have numbers. We will clean the data by removing entries with no IATA valid code. Here are the original entries:
![]()
We will retain cleaned-up rows in the total amount that follows:

Marco: Next, we create a list of rules for all airport IDs and their coordinates:

Vitaliy: We used the Dispatch function, which generates an optimized dispatch table representation of a list of rules and will never affect results that are obtained, but may make the application of long lists of rules much faster.
Marco: Now we can calculate the connections:

Not every IATA code has geo coordinates. Let’s clean out those with missing data:

Out of a total of 67,210, we will plot just 15,000 random routes, which reflects well on the full picture:
Vitaliy: Once we have the data, how can it be integrated with mathematical models?
Marco: We need to describe the mobility pattern of the population. We will use the global air transport network to build a first model of a pandemic. We think of the flights as connections between different areas. These areas could be interpreted as “catchment areas” of the respective airports.
Vitaliy: We could make use of the Graph function to generate the network quickly and efficiently. There are many different routes between the same airports, and this would correspond to multigraph, a new Wolfram Language feature. For the sake of simplicity of a starting model, we will consider only the fact of connection between two airports, drawing a single edge if there is at least one route. We will use multigraphs later in the improved model:
![]()
In the resulting graph, vertices are given by IATA codes.
As we can see below, there are several disconnected components that are negligible due to relatively small size. We will discard them, as they will not significantly alter any dynamics:

From our graph, we can construct the adjacency matrix:

Marco: The (i,j)th entry of the matrix is zero if the airports i and j are not connected, and one otherwise. The coupling matrix is automatically converted into the SparseArray form to make computations and memory use more efficient. We can also use a “population size” term for each airport, that is, its catchment area. We will later try to make more reasonable guesses at the population size, but for now we will simply assume that it is the same for each airport:
![]()
Now comes a tricky step. We need to define the parameters for our model. The problem is that each of them “parametrizes” many effects and situations. For our model, we need:
- The probability of infection. This is a factor that determines in our model how likely it is that a contact leads to an infection. This factor will certainly depend on many things: population density, local behavior of people, health education, and so on. To get our modeling started, we will choose the following for all airports:
![]()
- The rate of recovery. The rate of recovery will very much depend on the type of disease and the quality of the health system. It will also depend on whether everyone has access to health insurance. In countries where only a fraction of the population has access to high-quality health care, diseases will generally spread faster. For our initial modeling, we will set the following for all airports:
![]()
With these two parameters, we have the epidemic model determined. But we still need one more parameter.
- Migration factor. It is a factor of proportionality that describes the propensity of a certain population (in the catchment area of an airport) to travel. In this model, we have taken it to be constant, but it would certainly also depend on the financial situation in that country and other factors. It describes, roughly speaking, the percentage of people in a catchment area/country who travel. We do not use a multi-agent model where the movement of individuals is described. We use a compartmental population model where, in fact, we describe percentages of the population who travel. We do (at least later in the post, for the country-based simulation) take the different population size into consideration, and in the form of the multigraph, also how many flights there are from country to country. Using the coupling matrix, we will not introduce a migration of individuals between different airports. We will first use a general migration factor (same for all), which we set to the following:
![]()
This is a strong assumption. We will use—at first—the same migration factor for all three categories: susceptibles, infected, and recovered. In reality, the infected, particularly in the infectious phase where they might have developed symptoms, will probably have a different mobility pattern. Also, the mobility will be different in different countries, and it will depend on the distance traveled as well. We will later choose more realistic parameters.
We next initialize our model and assume that at first there are only susceptibles in all cities and no infected or recovered:
![]()
Now we should introduce 5% infected to the catchment area of the original airport.
Vitaliy: The outbreak began in Guinea in December 2013, then spread to Liberia and Sierra Leone. According to The New York Times, the very first case treated outside West Africa was a nurse from Spain infected in September while treating a missionary who had been infected in Sierra Leone. She was then flown to a hospital in Madrid. There are a few airports in Guinea:

We choose the CKY code for the Conakry International Airport in the capital and the largest city and compute its index:

Marco: Before we write down the SIR equations with the coupling terms, we introduce two objects:
![]()
This is the total number of (potential) passengers at each airport. We condense the coupling matrix into a smaller list that for each airport only contains a list of connected airports:
![]()
This is very useful, because the coupling matrix is sparse and sumind will speed up the calculation dramatically. Now we can write down the SIR equations:

The coupling terms are highlighted in orange. Basically, we calculate a weighted average over the number of people in each compartment for all neighboring airports. There are many other types of coupling that one could choose.
Next we iterate the equations and calculate the time course:

To get a first impression, we can plot the S, I, and R curves for some 200 airports:

Next, we calculate the maximal number of sick people at any of the airports:
![]()
We will generate a list of airport coordinates ordered as vertices of our graph:
![]()
We can now look at a couple of frames:
Time progresses by columns from top to bottom and between the columns from left to right. Here and below in a similar simulation graphic, the color codes the number of infected people.
Note that there are three main regions: Europe, which gets infected first, then the Americas and Asia. This kind of spreading is related to the network structure. We can try to visualize this with Graph.
Vitaliy: The clustering algorithms of Graph can help us see which continents are major facilitators in the spread of the pandemic via airways. We’ll make use of built-in data to get the list of continents:

We’ll discard Antarctica and build a database denoting which airport codes belong to which continent:
This is a function that can tell what continent a particular code belongs to:
![]()
For instance:

There are many differently sized communities in our network:

Communities are clusters of many flights joining airports of the same community compared to few flights joining airports of different communities. Not to overcrowd the plot with labels, let’s label only those communities whose size is greater than 60, based on the largest fraction of airport codes belonging to the labeling continent:

Now we can visualize the network structure and see how major hubs enable transportation to smaller ones. In this particular plot, colors just differentiate between different communities and do not represent infected people:
Marco: The three dominating communities become apparent. The graphic also shows via which of the main communities smaller groups are infected. This could have implications for decisions about preventative measures. We can now generate a graph that is similar to one presented in Dirk Brockmann’s paper:
Vitaliy: Again, time progresses by columns from top to bottom and between the columns from left to right. We used “RadialEmbedding” for GraphLayout and set “RootVertex” to airport. I see that the pandemic proceeds from the vertex that is central outward through the hierarchy layers. Marco, could you please explain what this means?
Marco: In the center of the representation, we find the airport where the outbreak started. The layer around it represents all the airports that can be reached with direct connections; they are the first to be hit. The next layer is the airports that can be reached with one connection flight; they are the next to be infected, and so on. This shows that the structure of the network, rather than geographical distance, is important.
Vitaliy: How can we make this model a bit more realistic?
Marco: We have studied some simple, prototypical cases of a model that locally describes the outbreak of a disease with an SIR model and then couples many SIR models based on connections of some transport network. However, there are many problems that still would have to be addressed:
- The probability of transmission will depend on many factors, for example, the health system and the population density in a region.
- The recovery rate will also depend on many factors, for example, the health system.
- Not all possible links (roads/flight trajectories) will be taken with equal probability. Published papers suggest that there is a power law: the longer the distance, the less likely someone is to travel.
- The migration/traveling rate will depend on the categories susceptibles, infected, recovered; sick people are less likely to travel.
Also we might want to be able to model different attempts by governments to control the outbreak. So let’s try to address some of these issues and generate another, global model of an Ebola outbreak. If we wanted to model all cities and all airports worldwide, that would probably be asking too much from an ordinary laptop. Based on the sections above, it should be clear how to extend the model, but I want to highlight some further approaches that might be useful.
Our model will represent all (or most) countries worldwide. The connections among them will be modeled based on the respective flight connections. To start with, we will collect some data for our model.
As before, we will import the airport data and the flight connections:

The SemanticImport function directly recognizes the countries that the airports belong to. We can easily construct a list of airports-to-country data:
![]()
This time we construct the graph using the Graph command. As we want to study the connections between countries, we substitute the airports by the respective countries:
![]()
Vitaliy: Are you saying that the graph connections indicate country adjacencies rather than airport connections?
Marco: The connections are generated by the flights; that is, the flights’ paths are the edges. They go from airport to airport, but we are actually interested in modeling the connections between countries. Therefore, we substitute (identify) the airports with their respective countries. Formally, you can think of this as constructing the network of all airport connections and then identifying all airports (nodes) that correspond to the same country. We could say that the mathematical term for that is vertex identification.
It turns out that data on some airports is missing; most of the missing data is from very small airports. We delete all airports for which the country is not known:
![]()
Last but not least, we construct the coupling/adjacency matrix:
![]()
We can also get the list of all countries that will form part of our model:
![]()
It is clear that if the population density is higher, that might lead to a higher infection rate; this is, of course, quite an assumption, because what is important is mostly local population densities. If the country is very large, but everybody lives in one city, the effective density is much higher.
To take the population into consideration, we will need data on the population size and density for all countries:

Now we will build a simple “model” for how the population density might (partially) determine the infection rate. In the first model we used an infection rate of ρ=0.2, and that gave “reasonable” results, in the sense that we saw a pattern of the spreading that was as expected. We want to extend the model, without completely modifying the parameter range. So it is “reasonable”—as a first guess—to choose the parameters for the extended model in the same range. As a matter of fact, we observe that the crucial thing is the ratio of λ and ρ. So basically we are saying that we want to start from about the same ratio as in the first simulation.
To modify the infection rate with respect to the population density, we look first at the histogram of the population densities:

We can also calculate the median:
![]()
We will make the assumption that the infection rate will increase with the population density. For the “median population density” we wish to obtain an infection rate of 0.2. We will make the bold assumption that the relationship is given by the following chart:

We can calculate the infection rates for all countries:
![]()
Of course, this “model” for the dependence of the infection rate on the population density is very crude. I invite the reader to improve that model!
Vitaliy: It would be really interesting to know some typical laws, based on real data, about how infection rate varies with population density. It is great to have this law as a changeable condition so that readers can try their versions of it and observe change in the simulations. Percolation phenomena perhaps should be considered, where there is a sudden steep change in infection rate when population density reaches some critical value. There could also be some saturation behavior due to a city’s infrastructure. But I am just guessing here. Would you please share your own take on this?
Marco: If there are more people in a confined space, the infection rate should increase monotonically. It also appears to make sense that the infection rate does not increase linearly, but that its derivative decreases (perhaps even saturates). The infection rate should somewhat depend on the distance to the next neighbor, that is, go with the square root of the density, which would lead to an exponent of 0.5. But then the movement of individual people would be blocked by their neighbors, effectively decreasing the slope, so we would need an exponent smaller than 0.5. Well that was at least my thought. The article “The Scaling of Contact Rates with Population Density for the Infectious Disease Models” by Hu et al., 2013, shows that our assumptions are reasonable—at least for this crude model.
To estimate the recovery rate, we will make another very daring assumption. We will assume that the health system is better in countries where the life expectancy is higher. To build this submodel, we will make use of the life expectancy data built into Mathematica:
![]()
We can visualize the distribution of life expectancies for all countries in a histogram:

Note that there are a few countries that are lacking this data. We set their life expectancies to a typical 70; this will not influence the results, because these countries (very small countries, i.e. islands, and Antarctica) will not play a role in our model.
The median life expectancy:
![]()
Now, like before, we will formulate our assumption that life expectancy is a proxy (approximate indicator) for the quality of the health system, which is a proxy for the recovery rate. This assumption is again quite crude, because cases in relatively rich countries like Spain and the US suggest that recovery rates might not be substantially higher in wealthy countries:

As with the infection rates, we calculate the recovery rates for all countries:
![]()
We can also represent what that means for all countries; the median values are marked in white and blue, while the parity of the infection rate and the recovery rate is indicated by the black dashed line. The background color represents the percentage of people who contract the disease at any time during the outbreak, as derived from the SIR model (see above):

Vitaliy: A typical modeling process entails running the simulation many times, trying to understand the system behavior and the influence of parameter values. Our particular values will result in some common-sense final outcome for what we can imagine will happen fighting a very tough but survivable pandemic. Countries with weak economies will suffer the highest damage, while their counterparts should count smaller losses. What is important here, I think, is to understand how that contrast depends on mobility network topology, demographics, and other real-world factors. We also should expect that these complex components and nonlinearity could result in counterintuitive behavior sometimes favoring the weak and damaging the strong.
Marco, could you please explain in greater detail the plot above?
Marco: The plot above shows the main two parameters of our model: infection versus recovery rate. Each point denotes a country—hovering over them will display the name. The white and green lines indicate the median values. They cut the diagram into four areas. The upper-left box (high infection rate/low recovery rate) contains countries that have the most challenging conditions. There are countries such as Sierra Leone, Nigeria, and Bangladesh in it. In the lower right (low infection rate/high recovery rate) are the countries that have the best prospects of containing the disease: United States, Canada, and Sweden. The black dashed line indicates a critical separator: above the line there are countries where the infection rate outpaces the recovery rate. In those countries an outbreak is very likely. Below the line the recovery rate dominates the infection rate, and the chances are that the disease can be contained. Note that these are just “base parameters”; they do not take the network into consideration yet. Note that countries below the dashed line can block the spread of the disease. If there are enough of them, that again will decrease the number of casualties.
Vitaliy: As we have mentioned before, our simulation is significantly exaggerated to amplify and see clearly various effects of a pandemic. This is why we intentionally shifted the system above the black dashed line. Our readers are welcome to consider more realistic parameter values.
Marco: To address another flaw of our first models, we will assume that susceptibles, infected, and recovered travel at different rates. There is one additional point to consider. If we set the migration/movement rates too high, regional differences will average out! If susceptibles, infected, and recovered all travel (fast enough), the populations of different countries mix over the periods of the simulation, so that we just see an average. In reality, the percentages of people traveling will be relatively low over the simulation period, so they will nearly be zero. In order to see the disease spreading, we will set relatively high rates for the movement of the infected; we will ignore the traveling of the healthy and recovered. This is not a bad assumption, because they do not contribute to the spreading of the disease anyway:
![]()
Let’s clear the variables to make sure that we start with a clean slate:
![]()
As before, we initiate all countries to have purely susceptible populations:
![]()
The recent outbreak started in Sierra Leone, so we will need to find out which vertex models that country:

Next we set a low number of infected people in that country:

We then iterate as before:

Now we can actually iterate:

This is a first test of whether the simulation worked—we plot time series for all countries:

Note that at the end of the simulation, most of the lines have saturated, indicating that the outbreak has come to a standstill. Here is also a time course of the total sick and deaths over the course of the outbreak:

We can now visualize the global spread of Ebola with the help of a few time frames:
Note that the number of dead people is extremely high in all of our simulations. About three billion dead people at the end would be an absolutely devastating outbreak, most likely much, much larger than what we are currently observing. This might be due to the choice of parameters; the ratio of the infection rate to the recovery rate might be different. On the other hand, we model an outbreak where the systems are relatively “inert,” meaning that they do not take efficient countermeasures. If we were to model that, we would need to decrease the infection rate over time, due to the actions of the governments.
The model is indeed just a conceptual model at this point. The reader can change certain features, that is, close down airports, and see what the effect is. He/she can change infection and recovery rates and see how that influences the outcome. There are many things you can play with. (A really bad infection rate/recovery rate ratio might be important if Ebola should mutate and become airborne, for example.) We have not (yet) tried to choose the parameters so that they optimally describe the current outbreak.
Vitaliy: I run our model many times for different parameter values. For example, in an opposite to the above situation, when recovery is greater than infection rate and the initial values are smaller:

We get much lower infected and sick numbers:

Also note different peak behavior, which tells us that pandemics can relapse—and with greater potential casualties. The New York Times data shows such relapses in the section “How Does This Compare to Past Outbreaks?” Also the C.D.C. projection and current stage of pandemic are probably an approximate exponential increase matching the very beginning (left side) of peaks in the graphs.
Can we calibrate our model by real data, for instance, time scale and absolute values of infection and recovery rates?
Marco: There are many papers that study effects of, say, population density on infection rates, such as the one mentioned above by Hu et al. We can use models or observed data to calibrate our model. Mathematica offers many great tools to fit parametric curves (e.g. from models) to data. This can help to improve our model. Regarding the time scale, we need to stress that we have not yet attached a time unit to each iteration step. We can, however, use observed data, for example, for the current outbreak, and try to find parameters ρ, λ, and μ so that the number of casualties/infected and its change over time are correctly described by the model.
Vitaliy: The ListLinePlot and GeoRegionValuePlot both show behavior different from the simplified original model we started from. Now we have countries that get outbreaks and others that seem resistant to the pandemic. This looks more realistic to me. Could you please share your thoughts about this and perhaps compare these results to Dirk Brockmann’s paper?
Marco: The most important factor is that the high migration rates in the first models lead to a mixing of the populations so that at the end, all populations have the same rates of people in each of the three categories. Just like in Dirk Brockmann’s paper, we see that the network is very important for the spreading of the disease. The distance on the graph is important. This is particularly so because countries for which the recovery rate is higher than the infection rate basically block the disease from spreading.
Vitaliy: Marco, I thank you very much for your inspiring insights and time. Do you have some ideas how we could improve the model further or any other concluding remarks for our readers?
Marco: It is very easy to come up with more ideas about how to improve the model: we could include further transport mechanisms, that is, streets/cars, boats, train networks, etc. Each would have an effect on how people move. We could use mobile phone data or other data to better model how people actually move. Also, the SIR model does not take into consideration the incubation time, that is, the time from infection until you show the first symptoms. Our model describes populations in compartments; numbers of infected etc. are given in percentages. This is only a valid approximation if the number of infected is large enough (like a concentration given as a real number is a good idea if there are lots of molecules!), but particularly at the beginning of an outbreak, when there are few infected, other types of model such as a multi-agent model might be more realistic. The model we have presented should only be taken as a first step; more effects can be included and their relevance can be studied. By trial and error, we can try to describe the real outbreak better and better. But we also need to be careful: the more effects we include in our model, the more parameters we will get. That can lead to all sorts of problems when we need to estimate them.
We may note that the simulation does not correctly describe the cases in the US and Europe. Currently we know that 18 people were brought there on purpose. They are not at all part of the natural propagation of the disease. Also, we are speaking about “percents or more” of a population, we are not even discussing individual cases. Epidemiologically speaking, there is nothing going on in the US or Spain at the moment regarding Ebola. Sierra Leone and other countries in Africa do get into a regime where the model is “more valid.”
The model is not specific to the Ebola epidemic. There is a lot of estimation and guesswork, and we choose the parameters so that the epidemic spreading becomes clear, but the model shows a potential order of the spreading: first mainly western/central African countries, then Europe, then the US. This is very reasonable and coincides with much more complicated models. Of course, there are many other effects, such as that the spreading between neighboring countries in Africa will probably not be via flights, but rather via very local transport, that is, people crossing borders by foot/car or similar. Our model is certainly conceptual in so far as it only considers one way of transport, namely flights, which is not the full picture, of course, but it does show that there are dramatic differences between countries to be expected. In fact, we would expect a much lower percentage of casualties in Europe and the US than in the countries in Africa were everything originated. The model shows that the highest probability of spreading is between neighboring African countries, which is what larger models predict as well. It also says that certain countries in Europe, such as Germany/UK/France etc. are more at risk than others because of their flight connections. The US would be less at risk than the European countries, that is, it would get significant numbers of infected later. All of that seems to be qualitatively quite correct. Australia and Greenland would get the disease very late, or not at all, again in agreement with our model (well, at least if rho ~ lambda ~ 0.1).
In that sense I suppose that it is more realistic to chose rho equal or smaller than lambda. That will mostly limit the spreading to some African countries with low probabilities of infections in Europe and the US. Even though the “global average” infection rate could be even lower than the recovery rate, at least in some countries, due to population density and (lack of) quality of health systems, the local infection rate might be higher than the recovery rate. Also, the number of infected will obviously be lower than what our model predicts, as our model does not take countermeasures by governments or the WHO into consideration. That is easy to model by reducing the infection rate over time, more in rich countries, less in poor countries.
The predictions have to be seen as probabilities or risk that certain countries experience large natural outbreaks, that is, our model does not consider individual cases or cases that are transported to hospitals on purpose. Also, the infection rate is certainly very different for people who work in hospitals with patients. That means that the probability for a nurse contracting Ebola is much higher than for the average person, and of course we do not model that. Also the precise propagation of an epidemic depends on many “random” factors, which cannot be reasonably taken into account in this or any other model. One can introduce random factors and run the model various times to make predictions about likely scenarios, or probabilities. In that sense, our model performs actually qualitatively very well.
The conceptual nature of the model allows us to look at different scenarios: What if infection rate versus recovery rate changes? What if there is more mobility, that is, mu changes? What if we also use local transport etc.? We did not try to fit the parameters/network etc. to optimally describe the Ebola outbreak; rather, we provide the basic model to develop several scenarios and to understand the basic ingredients for this type of modeling.
Vitaliy: I once again thank Marco and our readers for being part of this interview and invite everyone to participate in the related Wolfram Community thread where Marco and others discuss and share related ideas, code, and in-depth studies.








The mortality rate would be complex to calculate because of the changing status of the medical services in each area as they become overwhelmed, or not. I expect China’s ability to brutally control it’s population will, as it has in the past, make a big difference to the rate of infection whereas more open or more chaotic societies may fail to impose the necessary control over their populations, giving the same result but for different reasons.
There are also regional difference in the actual outbreak that need explaining if the spread is to be understood well enough to be modelled, take Côte d’Ivoire for example. If you look at the numbers in the effected countries at a finer grained level you see hot spots and gradients, yet the primary hotspot’s gradient to the east is truncated abruptly at the border with Côte d’Ivoire. What variable is influencing that?
I dare to send you my recent paper. I think that it can modify the complexity level that you considered in your model. The reference is “On evaluation of infection probability for different age groups” BioSystems, 112 82013 pp.305-308. Unfotunately I can not send the paper in the framework of the comment but if you want I can send it on your e-mail address.
Thank you! We’d love to hear more of your ideas. I think the best place for a technical discussion is our Wolfram Community – you can also attach files there if you wish to do so for your paper: http://wolfr.am/1BB1_ZVL
nice post Vitaliy!
Great article Vitaliy!
How is the death rate determined from the recovery rate for each country?
Thank you, Michael! We simply consider that 50% of total recovered are fatal cases, which is a conventional assumption in such models and is reasonable for average statistics over various countries in, for example, Ebola case. You can see this 50% in Out[90] – as the factor 0.5*Total[…]
You should sell this to a Hollywood script writer. If I remember correctly Michael Trott worked on Numbers so you might still have the contacts. In my opinion, you should add another twitch: The impact of trying to treat the disease. In many if not most cases, Ebola outbreaks were not discovered until it had eradicated almost the entire village. The virus would not spread since the infected could not leave the close vicinity. So how do you model a treatment that involves moving people around and risking that those who (mis)treat them in hospitals further away are not perfectly efficient?
Thank you, Ernst, you picked up quite precisely on some parallels to NUMB3RS here, – Marco and I talked about it a few times. In relation to your comment “…you should add another twitch: The impact of trying to treat the disease” I should point to Marco’s thoughts just below Out[95]: “…we model an outbreak where the systems are relatively “inert,” meaning that they do not take efficient countermeasures. If we were to model that, we would need to decrease the infection rate over time, due to the actions of the governments.” But you seem to pose here a more detailed problem in your last sentence. Generally, I think, any type of corrections should be introduced as an additional mobility layer represented as network, perhaps with dynamically weighted edges and vertices to model the system most lithely. For example, as a next iteration, we were considering a simplistic ground transportation layer added, as a Delaunay triangulation rooted in all cities, which we will probably add as an example on the original Wolfram Community discussion: http://wolfr.am/1BB1_ZVL
The use of geospatial data and the transport network is good, but the underlying SIR is way too primitive to take seriously Marko’s predictions.
The spread of Ebola in West Africa is first and foremost the result of inadequate (non-existent) primary healthcare system. Unlike respiratory (air-borne) infections (flu, SARS) ebola is highly inefficient in human spread (BRN close or less than 1).
Any serious attempt at modeling its spread should account for (i) social connectivity and behavior (ii) environment and interventions (e.g. quarantine). Population level SIR is a far cry.
Model of these type are more appropriate for simulating Ebola panic rather that Ebola infection.
I agree that the SIR model is quite primitive. It has, however, been quite successfully used to model very homogeneous populations.The main model assumption is that if the people in each population behave in a similar way their social connectivity can be “parametrised” by the parameter for the infection rate. It is quite clear that this is not quite true globally, but I hope that the model suggests one way of taking this into consideration: by including a network of sub-populations which can all behave differently. By using country wide data, we still model at a rather coarse level, but it should be clear how to go forward from here. We could model the cities/regions in each country by individual SIR models and adapt the parameters so that they reflect the local behaviour. We could then go to even smaller sub-populations and so on. There are two main problems with that approach: (i) one is that we could get more and more equations and the required CPU time and memory would very soon require very large computers.(ii) the second is that for each additional layer of sub-populations we would dramatically increase the number of parameters. We could try to determine them from data, such as mobile phone data (http://www.technologyreview.com/news/530296/cell-phone-data-might-help-predict-ebolas-spread/) and other data sources, but more parameters do not necessarily improve the predictions of the model.
I think that the two points you raise at the beginning (that Ebola spreads very inefficiently in human populations and the importance of the health care systems) are not problematic for the model we present. The low infection rate can easily be modelled by choosing the corresponding parameter to be small. The importance of the health care system is actually used in the model. The problem was how to parametrise this… We used the life expectancy as a primitive proxy for the quality of the health care system. This is of course only a crude guess, but as you say it is very relevant for how the disease spreads and your feeling that this is important is reflected by our analysis.
I also totally agree that we do not model interventions (e.g. quarantine), and that certainly is very relevant. There is of course a relatively easy way to model this, which is to decrease the infection rate over time, as the countries take counter-measures. In countries that invest more into the fight of ebola the infection rates could be assumed to decrease faster. It only takes a line of code to take this into consideration, and I have simulations for that. (They were too detailed for this blog, but if you are interested I can post them on the Community.)
Finally, I think that all you say about the social connectivity, the low infection rates, interventions etc. can be modelled – at various degrees of granularity – if you tell me more about which assumptions you want to implement. If you have a better idea of how people move, e.g. you have telephone data, or use “Where is George” (http://www.wheresgeorge.com) we can implement your assumptions into the model, just as we did with the life expectancy and the population density. It is fun to add new processes and see how the model reacts. I hoped to introduce a model that is flexible enough for everyone to include their own ideas and test what impact they might have.
Also note that this is supposed to be a conceptual model. It certainly has limitations but it does show some reasonable features, e.g. France is at a higher risk of infection than many other European countries. This makes sense as there are many links (including flights) to former colonies in Africa. It is also reasonable that Europe gets infected before the United states and Asia… So the model does make some reasonable predictions.
I’d love to discuss that with you in the Community…
Cheers,
Marco
Wow this is very informative to see it laid out like this! It’s important to have some of these graphics in order to see the full scope of the epidemic. I hope we continue to see more resources like this. I have also been following updates on the Ebola Virus from many resources such as: http://www.metrex.com/news/ebola-virus
Hope we can bring more resources and visibility to Ebola!
You listed North America twice when generating Vitality data.
Excellent! It’s a good model about ebola_virus spread to reference. There’re some problems for me to resolve. One of the most important problems is the data, such as the “airports.dat” and “route.dat”. Could I have a look the data_source about the “airports.dat”?
At the end of the blog there is a download link to all files including “airports.dat” and “route.dat”. In the blog text when the data are introduced there is a discussion and link to the data webpage. If you have any technical questions could you please post them on our related forum discussion: http://community.wolfram.com/groups/-/m/t/326240
Mathematical modeling, through use of a powerful computer program, of the spread of an infectious disease can not only save a lot of human lives but can even prevent future epidemics and pandemics.
Congratulations, outstanding report !
Could you please tell me where you received the real data from about the initial number of infected that equals 5% of the population?